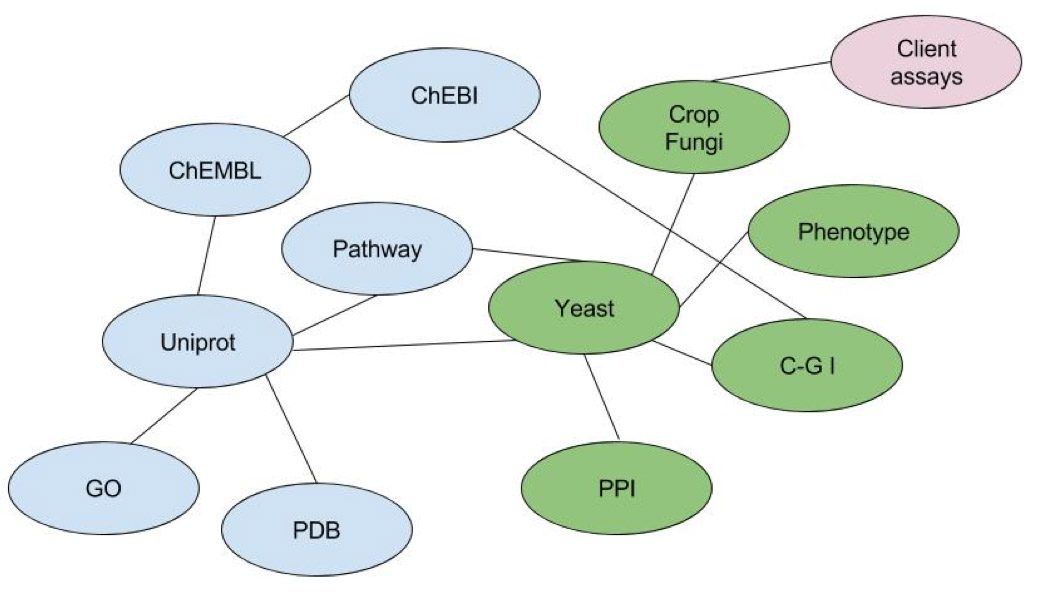
The scientific goal was to propose new targets and compounds for a fungicide chemical discovery pipeline and to validate the MOA of compounds currently in the pipeline. The project was triggered by a seminal publication of high-throughput yeast experimental data that was not tractable to scientists. We integrated that data and public annotation sources with client screening data, delivering new candidate compounds and targets and elucidated previously unknown modes of action. The legacy is a state of the art internal platform for target discovery from which the business continues to extract value.
Data Integration
The data required to meet the scientific requirements was
diverse and disparate. The data was heavily processed to allow
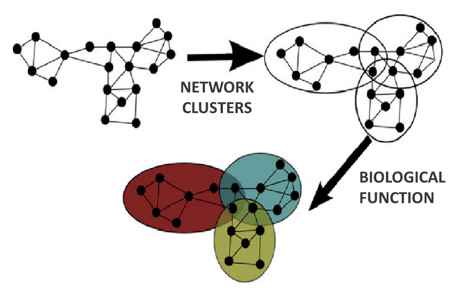
meaningful integration and analysis. The high-throughput
screening data was co-clustered to reveal interaction units
which were annotated with biological function. We used semantic
web methods to integrate all of the data and used Cytoscape to
interrogate networks.
The data continues to be updated and has been used in many ways with significant short and long term business impact, the client continues to use the data for proprietary target selection.
As an example of the power of integrated data we identified specific vs off-target effects for a battery of screening compounds. In numerous screens including tunicamycin only three genes were associated with the compound in all screens. The known target of tunicamycin was correctly identified along with two other genes with related biological roles which could be priority candidates for ongoing research.